Cap analysis of gene expression (CAGE) sequencing reveals alternative promoter usage in complex disease | bioRxiv
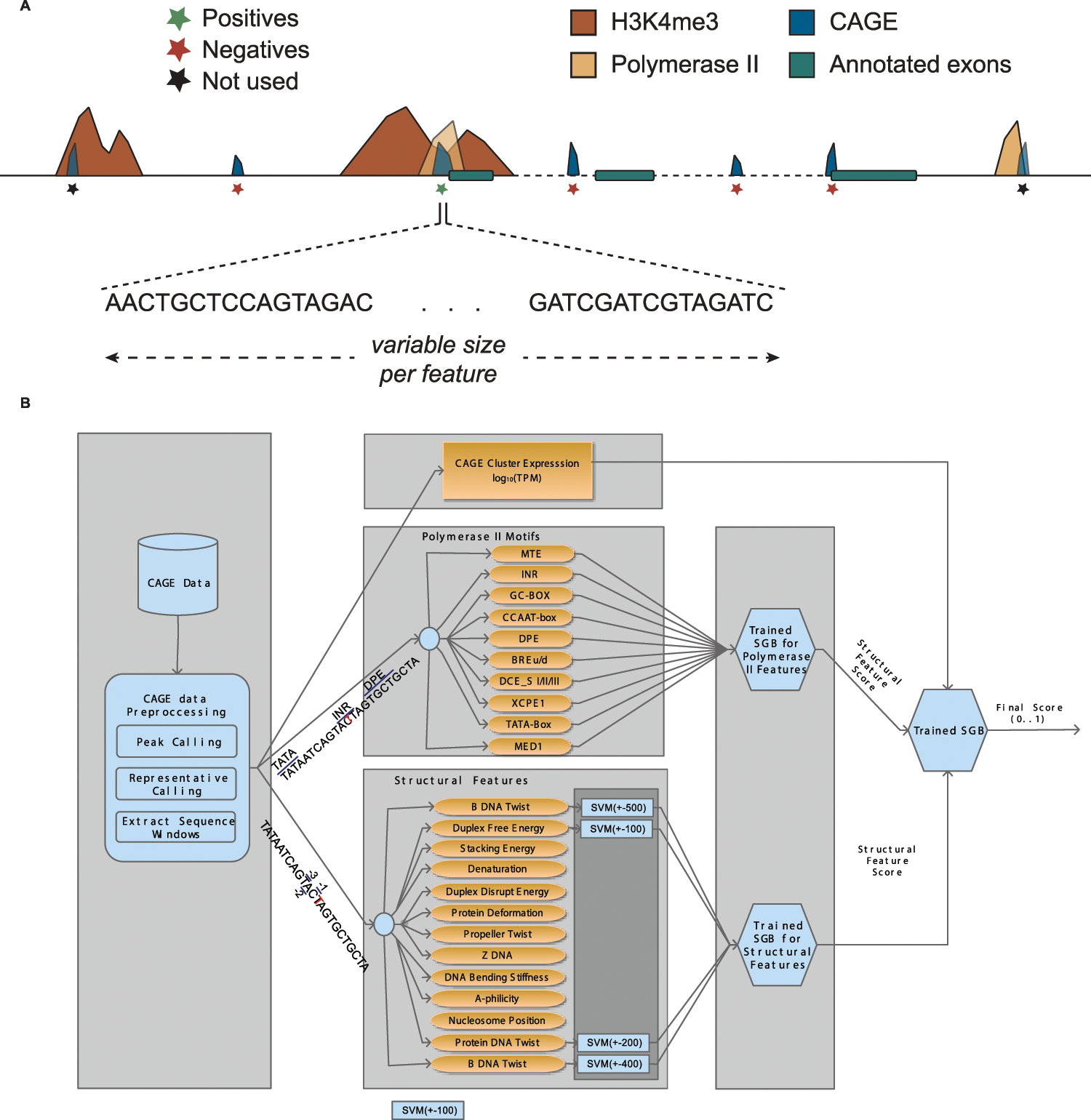
Solving the transcription start site identification problem with ADAPT-CAGE: a Machine Learning algorithm for the analysis of CAGE data | Scientific Reports

CAGE-seq characterization of alternative promoter usage and full-length... | Download Scientific Diagram
Comparison of CAGE-seq and RNA-seq gene expression levels. A. Scatter... | Download Scientific Diagram
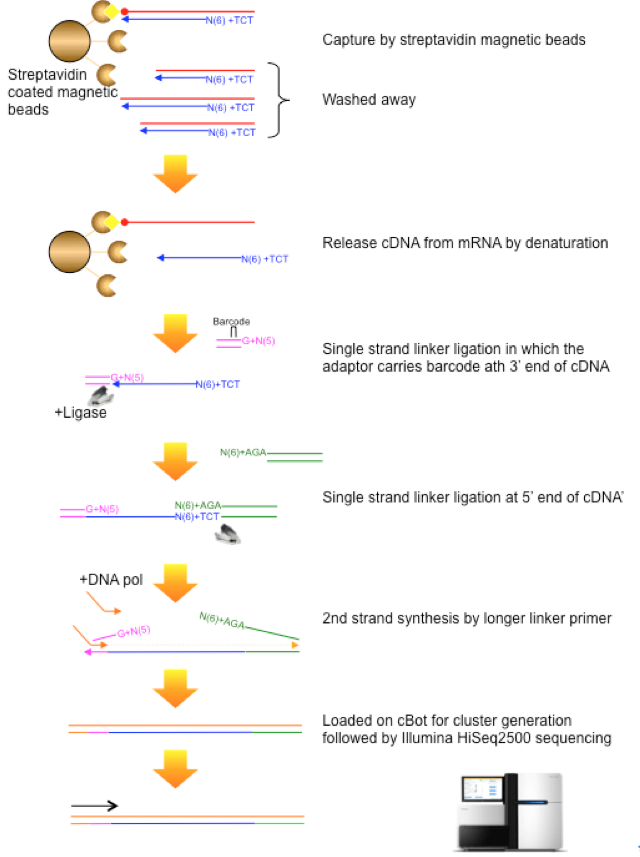
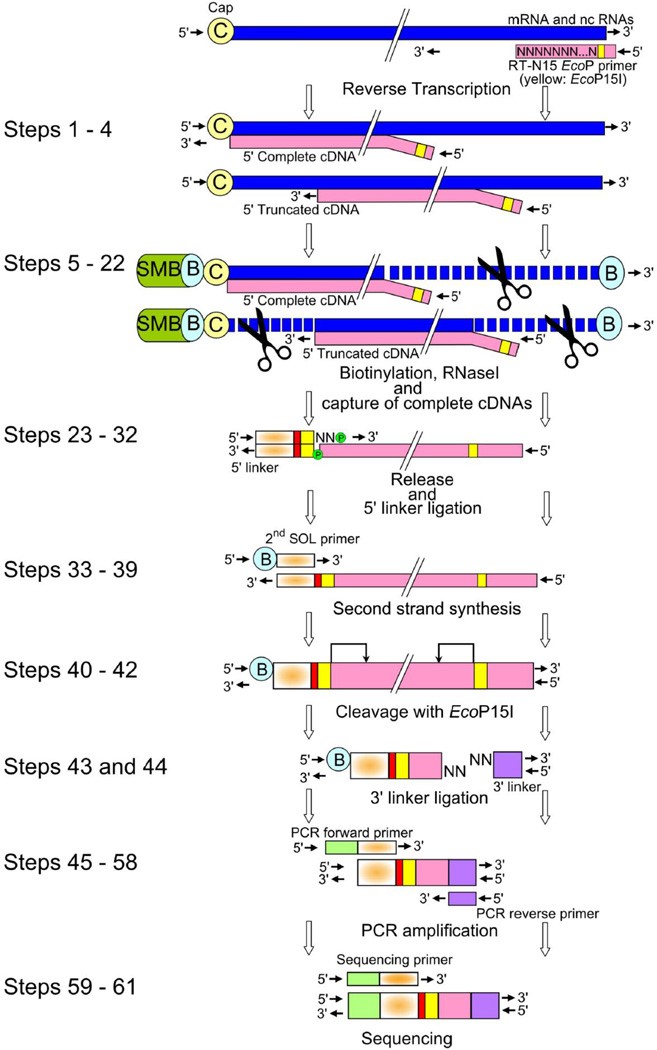
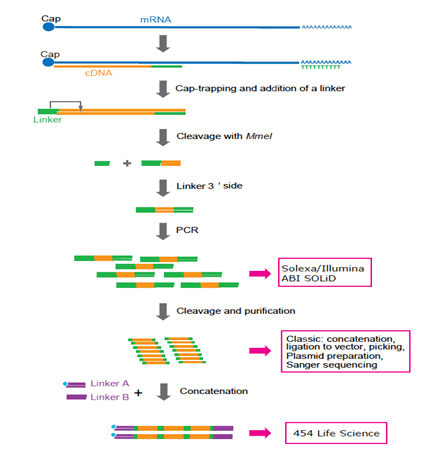
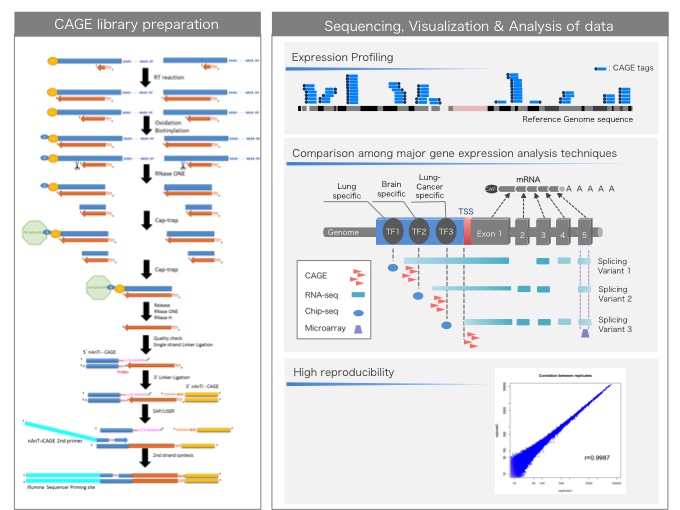
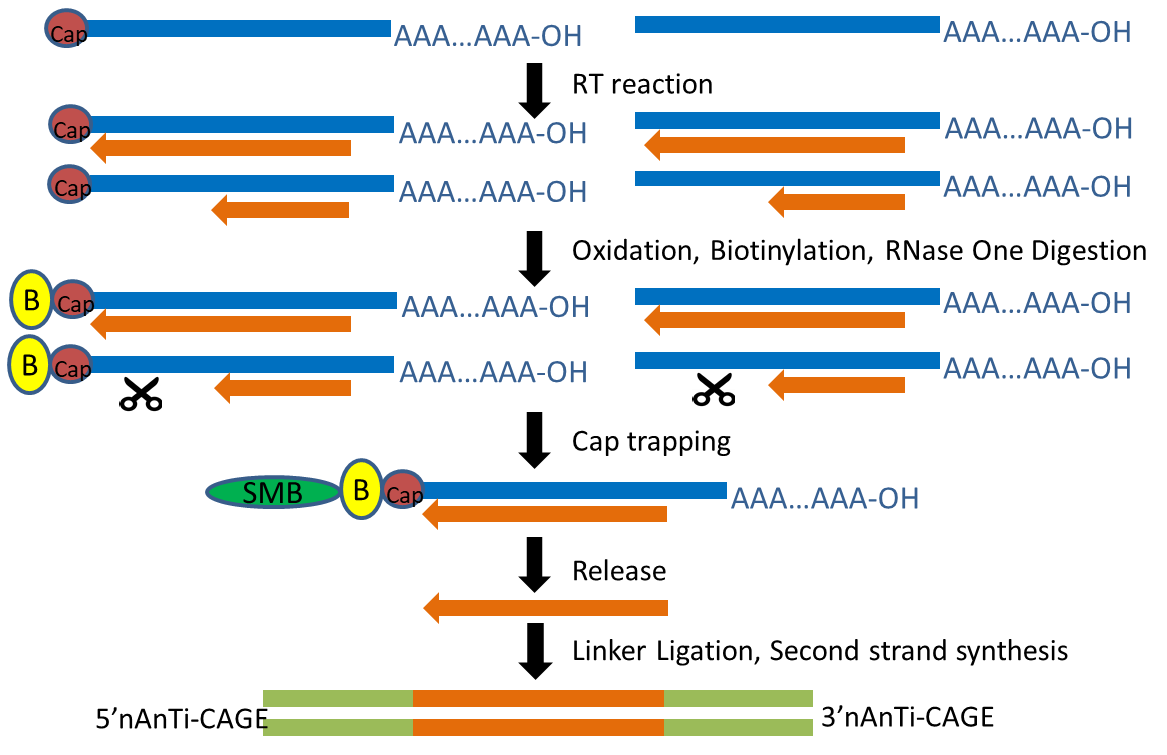
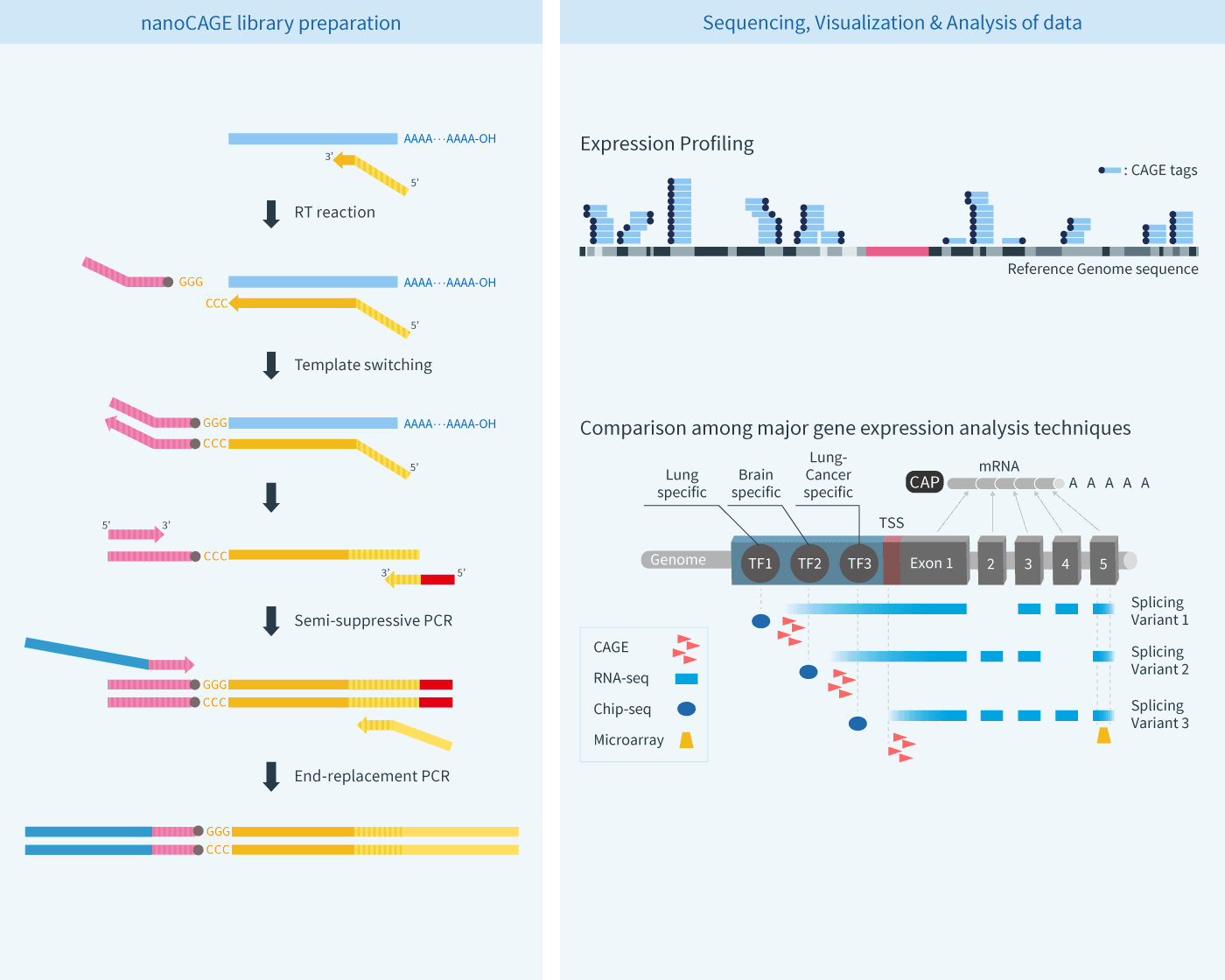
CAGE : A method for genome-wide identification of transcription start sites | 理化学研究所 ライフサイエンス技術基盤研究センター

Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv

Frontiers | Extensive Variation in Gene Expression is Revealed in 13 Fertility-Related Genes Using RNA-Seq, ISO-Seq, and CAGE-Seq From Brahman Cattle
CAGE-seq analysis of Epstein-Barr virus lytic gene transcription: 3 kinetic classes from 2 mechanisms | PLOS Pathogens

Integration of Single-Cell RNA- and CAGE-seq Reveals Tooth-Enriched Genes - Y. Chiba, K. Yoshizaki, T. Tian, K. Miyazaki, D. Martin, , Genomics and Computational Biology Core, Genomics and Computational Biology Core, K.

Cap analysis of gene expression reveals alternative promoter usage in a rat model of hypertension | Life Science Alliance